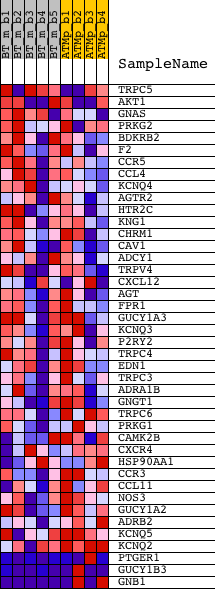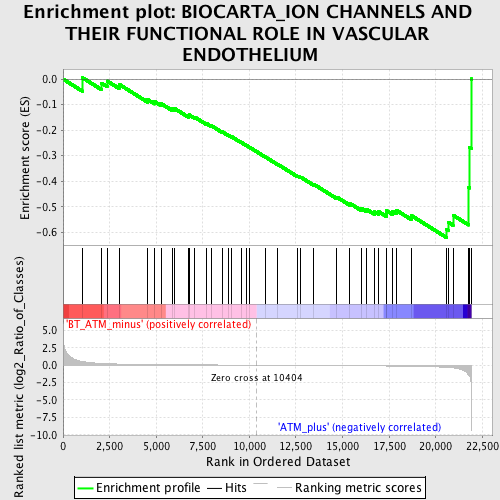
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
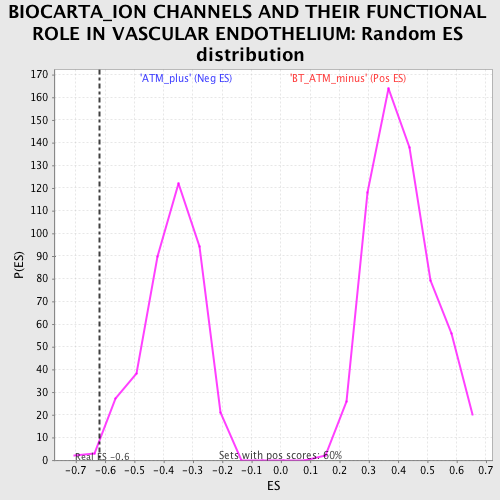
| GeneSet | BIOCARTA_ION CHANNELS AND THEIR FUNCTIONAL ROLE IN VASCULAR ENDOTHELIUM |
| Enrichment Score (ES) | -0.6204241 |
| Normalized Enrichment Score (NES) | -1.6652585 |
| Nominal p-value | 0.007556675 |
| FDR q-value | 0.25721976 |
| FWER p-Value | 0.989 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TRPC5 | 1422048_at 1442589_at 1459367_at | 1027 | 0.522 | 0.0061 | No | ||
| 2 | AKT1 | 1416657_at 1425711_a_at 1440950_at 1442759_at | 2060 | 0.252 | -0.0155 | No | ||
| 3 | GNAS | 1421740_at 1427789_s_at 1443007_at 1443375_at 1444767_at 1450186_s_at 1453413_at | 2363 | 0.216 | -0.0073 | No | ||
| 4 | PRKG2 | 1421354_at 1435162_at 1435460_at | 3007 | 0.161 | -0.0203 | No | ||
| 5 | BDKRB2 | 1422263_at 1442187_at | 4540 | 0.099 | -0.0802 | No | ||
| 6 | F2 | 1418897_at | 4917 | 0.089 | -0.0884 | No | ||
| 7 | CCR5 | 1422259_a_at 1422260_x_at 1424727_at | 5269 | 0.081 | -0.0962 | No | ||
| 8 | CCL4 | 1421578_at | 5852 | 0.069 | -0.1158 | No | ||
| 9 | KCNQ4 | 1435721_at 1442909_at | 5998 | 0.066 | -0.1157 | No | ||
| 10 | AGTR2 | 1415832_at | 6741 | 0.053 | -0.1442 | No | ||
| 11 | HTR2C | 1435513_at 1450477_at | 6766 | 0.052 | -0.1400 | No | ||
| 12 | KNG1 | 1416676_at 1426045_at | 7071 | 0.047 | -0.1491 | No | ||
| 13 | CHRM1 | 1439611_at 1450833_at | 7689 | 0.037 | -0.1735 | No | ||
| 14 | CAV1 | 1449145_a_at | 7955 | 0.033 | -0.1822 | No | ||
| 15 | ADCY1 | 1445112_at 1445359_at 1456487_at 1457251_x_at 1457329_at | 8534 | 0.025 | -0.2060 | No | ||
| 16 | TRPV4 | 1417545_at | 8878 | 0.020 | -0.2196 | No | ||
| 17 | CXCL12 | 1417574_at 1439084_at 1448823_at | 9049 | 0.018 | -0.2255 | No | ||
| 18 | AGT | 1423396_at | 9556 | 0.011 | -0.2475 | No | ||
| 19 | FPR1 | 1450808_at | 9828 | 0.008 | -0.2591 | No | ||
| 20 | GUCY1A3 | 1420533_at 1420534_at 1434141_at | 10007 | 0.005 | -0.2667 | No | ||
| 21 | KCNQ3 | 1458421_at | 10872 | -0.006 | -0.3055 | No | ||
| 22 | P2RY2 | 1450318_a_at | 11525 | -0.015 | -0.3338 | No | ||
| 23 | TRPC4 | 1423579_a_at 1446366_at 1451033_a_at | 12604 | -0.029 | -0.3801 | No | ||
| 24 | EDN1 | 1451924_a_at | 12748 | -0.031 | -0.3836 | No | ||
| 25 | TRPC3 | 1417577_at | 13471 | -0.041 | -0.4124 | No | ||
| 26 | ADRA1B | 1422183_a_at | 14696 | -0.060 | -0.4622 | No | ||
| 27 | GNGT1 | 1425167_a_at 1425168_at 1425219_x_at 1451633_a_at | 15403 | -0.072 | -0.4871 | No | ||
| 28 | TRPC6 | 1449431_at | 16026 | -0.083 | -0.5071 | No | ||
| 29 | PRKG1 | 1443663_at 1443811_at 1443812_x_at 1444232_at 1445807_at 1445998_at 1449876_at | 16300 | -0.088 | -0.5106 | No | ||
| 30 | CAMK2B | 1448676_at 1455869_at | 16726 | -0.098 | -0.5200 | No | ||
| 31 | CXCR4 | 1448710_at | 16946 | -0.103 | -0.5195 | No | ||
| 32 | HSP90AA1 | 1426645_at 1437497_a_at 1438902_a_at 1455078_at | 17352 | -0.114 | -0.5264 | No | ||
| 33 | CCR3 | 1422957_at | 17364 | -0.114 | -0.5153 | No | ||
| 34 | CCL11 | 1417789_at | 17688 | -0.123 | -0.5175 | No | ||
| 35 | NOS3 | 1422622_at | 17896 | -0.130 | -0.5138 | No | ||
| 36 | GUCY1A2 | 1459336_at | 18693 | -0.156 | -0.5343 | No | ||
| 37 | ADRB2 | 1437302_at | 20580 | -0.309 | -0.5890 | Yes | ||
| 38 | KCNQ5 | 1427426_at 1441071_at | 20679 | -0.325 | -0.5604 | Yes | ||
| 39 | KCNQ2 | 1420800_a_at 1426006_at 1426080_a_at 1426128_a_at 1426335_at 1435942_at 1438260_at 1440141_at 1440258_at 1451595_a_at 1459826_at | 20942 | -0.385 | -0.5332 | Yes | ||
| 40 | PTGER1 | 1436542_at 1445445_s_at 1450491_at | 21785 | -1.439 | -0.4255 | Yes | ||
| 41 | GUCY1B3 | 1420871_at 1420872_at | 21814 | -1.582 | -0.2660 | Yes | ||
| 42 | GNB1 | 1417432_a_at 1425908_at 1454696_at 1459442_at | 21915 | -2.671 | 0.0009 | Yes |